Interactive Volume-Based Visualization and Exploration for Diffusion Fiber Tracking
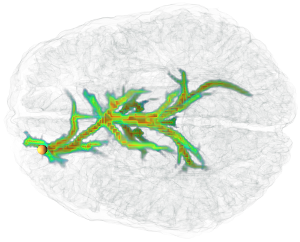
We present a new method to interactively compute and visualize fiber bundles extracted from a diffusion magnetic resonance image. It uses Dijkstra's shortest path algorithm to find globally optimal pathways from a given seed to all other voxels. Our distance function enables Dijkstra to generalize to larger voxel neighborhoods, resulting in fewer quantization artifacts of the orientations, while the shortest paths are still efficiently computable. Our volumetric fiber representation enables the usage of volume rendering techniques. Therefore no complicated pruning or analysis of the resulting fiber tree is needed in order to visualize important fibers. In fact, this can efficiently be done by changing a transfer function. Our application is highly interactive, allowing the user to focus completely on the exploration of the data.

