Publications
Extracting consistent and manifold interfaces from multi-valued volume data sets
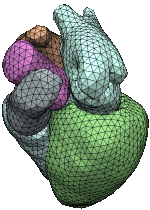
We propose an algorithm to construct a set of interfaces that separate the connected components of a multi-valued volume dataset. While each single interface is a manifold triangle mesh, two or more interfaces may join consistently along their common boundaries, i.e. there are no T-junctions or gaps. In contrast to previous work, our algorithm classifies and removes the topological ambiguities from the volume before extracting the interfaces. This not only allows for a simple and stable extraction algorithm, but also makes it possible to include user constraints.
Sub-Voxel Topology Control for Level Set Surfaces
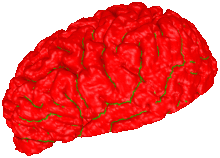
Active contour models are an efficient, accurate, and robust tool for the segmentation of 2D and 3D image data. In particular, geometric deformable models (GDM) that represent an active contour as the level set of an implicit function have proven to be very effective. GDMs, however, do not provide any topology control, i.e. contours may merge or split arbitrarily and hence change the genus of the reconstructed surface. This behavior is inadequate in settings like the segmentation of organic tissue or other objects whose genus is known beforehand. In this paper we describe a novel method to overcome this limitation while still preserving the favorable properties of the GDM setup. We achieve this by adding (sparse) topological information to the volume representation at locations where it is necessary to locally resolve topological ambiguities. Since the sparse topology information is attached to the edges of the voxel grid, we can reconstruct the interfaces where the deformable surface touches itself at sub-voxel accuracy. We also demonstrate the efficiency and robustness of our method on synthetic as well as on real (MRT) scan data.

